|
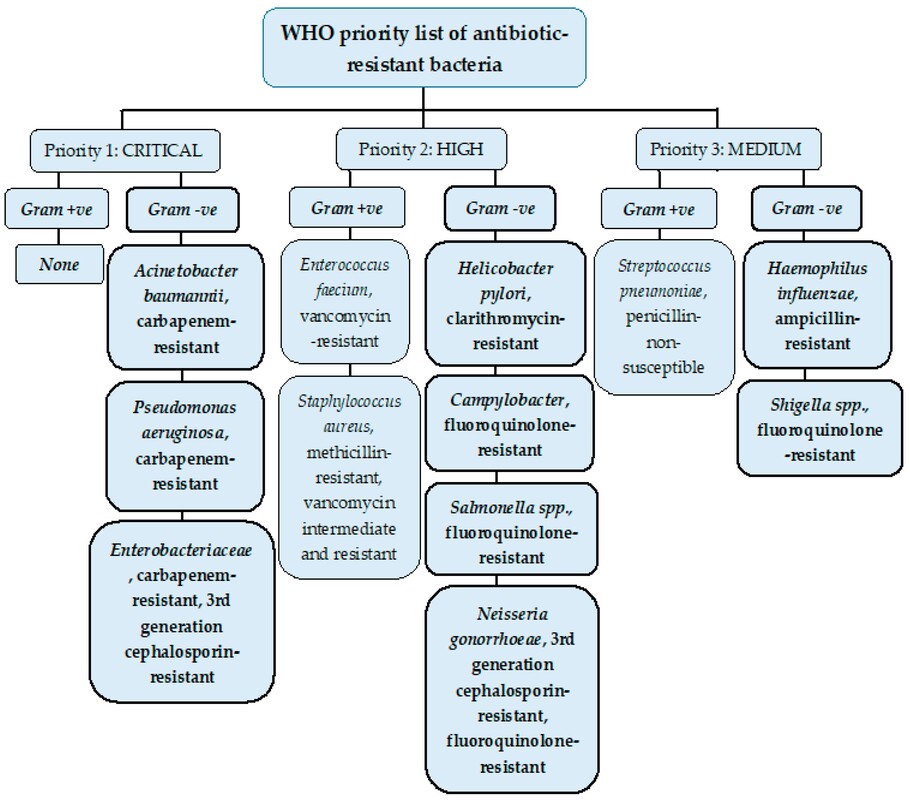
John writes … A century (nearly) of antibiotics It is one of those things that ‘every school pupil knows’: Alexander Fleming discovered penicillin in 1928. At the time, he thought that his discovery had no practical application. However, in 1939, a team at Oxford, led by Howard Florey, started to work on purification and storage of penicillin and having done that, conducted trials on animals followed by clinical trials with human patients. The work led to its use amongst Allied troops in the World War II and, after the war, in more general medical practice. Its detailed structure was worked out by Dorothy Hodgkin, also at Oxford, in 1945 and that led to the development of a range of synthetic modified penicillins which are more effective than the natural molecule. This leads us to think about penicillin’s mode of action: it works mainly by disrupting the synthesis of the bacterial cell wall which may in turn lead to the autolysis (self-destruction) of the cell. Other antibiotics have since been developed which target other aspects of bacterial metabolism. In my own research on genes, I have used rifampicin which inhibits the transcription (copying) of genes into messenger RNA (the working copy of a gene) and chloramphenicol which prevents the use of messenger RNA in the synthesis of proteins (1). However, antibiotics in the penicillin family are by far the most widely used (but see next section).  Hans Christian Gram (1853 -1938) Photograph by Niels Christian Hansen and Frantz Clemens Stephan Weller via Wikimedia Commons. Hans Christian Gram (1853 -1938) Photograph by Niels Christian Hansen and Frantz Clemens Stephan Weller via Wikimedia Commons. But there’s a problem Bacteria can be grouped according to whether they are ‘Gram-positive’ or ‘Gram-negative’. Hans Christian Gram was a Danish bacteriologist who developed a technique for staining bacterial cells – the Gram stain. Species which retain the stain, giving them a purple colour, are Gram-positive; species which do not retain the stain are Gram-negative. We now know that this difference is caused by differences in the cell’s outer layers. Gram-positive bacteria have a double-layered cell membrane (the plasma-membrane), surrounded by a thick cell wall; Gram-negative bacteria also have a double-layered plasma-membrane which is surrounded by a thinner cell wall and then another double-layered membrane. It is this structure that prevents the stain from entering the cells and which also leads to antibiotics in the penicillin family being totally or partially ineffective. Like many molecular biologists, I have used non-pathogenic strains of a Gram-negative bacterium, Escherichia coli (E .coli). It is the bacterial model used in research and has also been used to ‘look after’ genes which were destined for use in genetic modification. By contrast, pathogenic strains of this species can cause diarrhoea (‘coli’ indicates one of its habitats – the colon) while more seriously, some Gram-negative bacteria may cause pneumonia and sepsis. As already indicated, penicillin derivatives are mostly ineffective against Gram-negative bacteria and some of the effective antibiotics which have been developed have quite serious side effects. And an even bigger problem It is a truth universally acknowledged that an organism better adapted to an environment will be more successful than one that is less well adapted. This is of course a slightly ‘Austenesque’ way of talking about natural selection. Suppose then, that antibiotics are so widely used in medical, veterinary, or agricultural settings that they effectively become part of the environment in which bacteria live. Initially there will be a small number of bacteria that, for a number of reasons, are resistant to antibiotics. For example, some are able to de-activate a particular antibiotic and some can block the uptake of an antibiotic. Whatever the reason for the resistance, these resistant bacteria will clearly do better than the non-resistant members of the same species. The resistance genes will be passed on in cell division and the resistant forms will come to dominate the population, especially in locations and setting where antibiotics are widely used. And that, dear reader, is exactly what has happened. Antibiotic resistance is now widespread especially in Gram-negative, disease-causing bacteria, as is seen in the WHO priority list of 12 bacterial species/groups about which there is concern: nine of these are Gram-negative. The situation, already an emergency, is now regarded as critical in respect of three Gram-negative bacteria. Thinking about this from a Christian perspective, we might say that antibiotics are gifts available from God’s creation but humankind has not used those gifts wisely. But there is hope The growing awareness of resistance has catalysed a greatly increased effort in searching for new antibiotics. There are many thousands of organisms ‘out there’ remaining to be discovered, as is evident from a recent announcement from Kew about newly described plant and fungal species. It is equally likely that there is naturally occurring therapeutic chemicals, including antibiotics, also remaining to be discovered, perhaps even in recently described species (remembering that penicillin is synthesised by a fungus). There are also thousands of candidate-compounds that can be made in the lab. Concerted, systematic, computer-aided high throughput searches are beginning to yield results and several promising compounds have been found (2). Nevertheless, no new antibiotics that are effective against Gram-negative bacteria have been brought into medical or veterinary practice for over 50 years. However, things may be about to change. On January 4th, a headline on the BBC website stated ‘New antibiotic compound very exciting, expert says’ (3). The Guardian newspaper followed this with ‘Scientists hail new antibiotic that can kill drug-resistant bacteria.’ (4) The antibiotic is called Zosurabalpin and it was discovered in a high throughput screening programme (as mentioned earlier) that evaluated the potential of a large number of synthetic (lab-manufactured) candidate-compounds. In chemical terminology Zosurabalpin is a tethered macrocyclic peptide, an interesting and quite complex molecule. However, its most exciting features are firstly its target organisms and secondly its mode of action. Both of these are mentioned in the news articles that I have referred to and there is a much fuller account in the original research paper in Nature (5). Referring back to the WHO chart, we can see that one of the ‘critical’ antibiotic-resistant bacteria is Carbapenem-resistant Acinetobacter baumannii (known colloquially as Crab). Carbapenem is one of the few antibiotics available for treating infections of Gram-negative bacteria but this bacterial species has evolved resistance against it (as described above). This means that it is very difficult to treat pneumonia or sepsis caused by Crab. The effectiveness of Zosurabalpin against this organism is indeed very exciting. Further, I find real beauty in its mode of action in that it targets a feature that makes a Gram-negative bacterium what it is. One of the key components of the outer membrane is a complex molecule called a lipopolysaccharide, built of sugars and fats. The antibiotic disrupts the transport of the lipopolysaccharide from the cell to the outer membrane which in turn leads to the death of the cell. 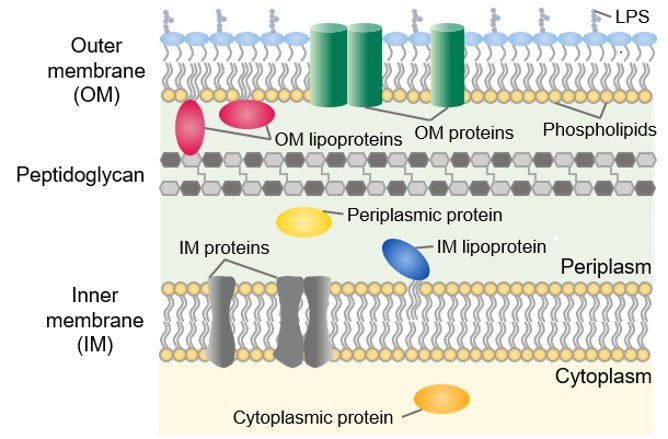 The outer layers of a Gram-negative bacterial cell in more detail, showing the inner membrane, the peptidoglycan cell wall and the outer membrane. The essential lipopolysaccharides mentioned in the text are labelled LPS. The new antibiotic targets the proteins that carry the LPS to their correct position. Diagram modified from original by European Molecular Biology Lab, Heidelberg. Zosurabalpin has been used, with a high level of success, to treat pneumonia and sepsis caused by A. baumannii in mice. Further, trials with healthy human subjects did not reveal any problematic side-effects. The next phase of evaluation will be Phase 1 clinical trials, the start of a long process before the new antibiotic can be brought into clinical practice. John Bryant Topsham, Devon January 2024 (1) For anyone interested in knowing more about how antibiotics work, there is a good description here: Action and resistance mechanisms of antibiotics: A Guide for Clinicians – PMC (nih.gov).
(2) As discussed here: Antibiotics in the clinical pipeline as of December 2022 | The Journal of Antibiotics (nature.com). (3) New antibiotic compound very exciting, expert says – BBC News. (4) Scientists hail new antibiotic that can kill drug-resistant bacteria | Infectious diseases | The Guardian. (5) A novel antibiotic class targeting the lipopolysaccharide transporter | Nature.
0 Comments
Leave a Reply. |
AuthorsJohn Bryant and Graham Swinerd comment on biology, physics and faith. Archives
July 2024
Categories |



 RSS Feed
RSS Feed
